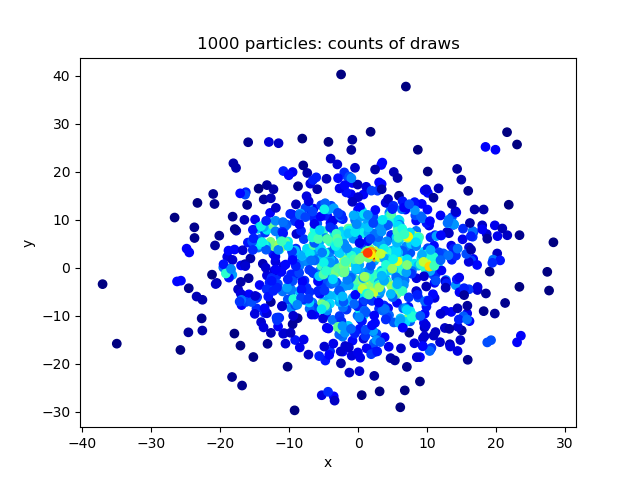
Physics-based machine learning for stochastic reaction networks
Methods and papers related to physics-based maching learning for modeling stochastic reaction networks, as developed in the thesis of Oliver K. Ernst
Research groups:
Eric Mjolsness Departments of Computer Science and Mathematics, UC Irvine
GitHub:
Click here to browse all code through this organization's GitHub page.
Thesis
Modeling reaction-diffusion systems with dynamic Boltzmann distributions
Papers & Code
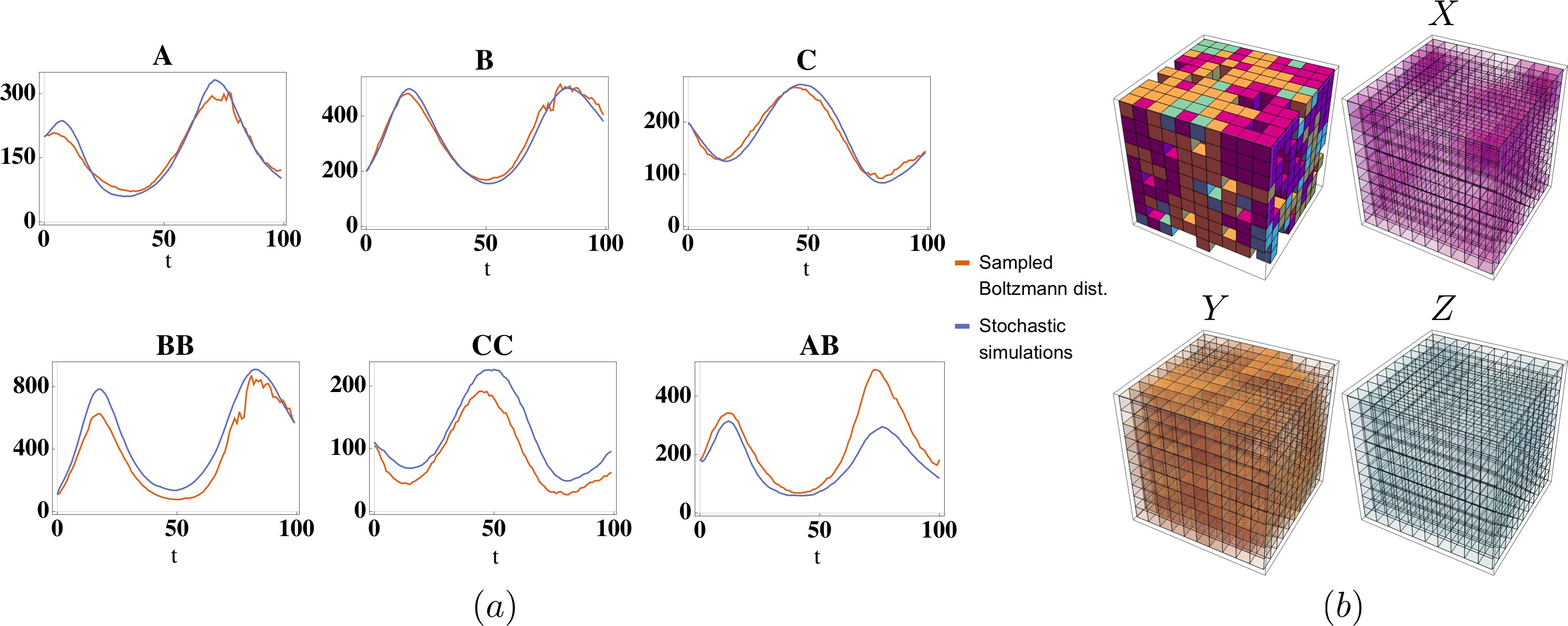
Physics-based machine learning for modeling stochastic IP3-dependent calcium dynamics
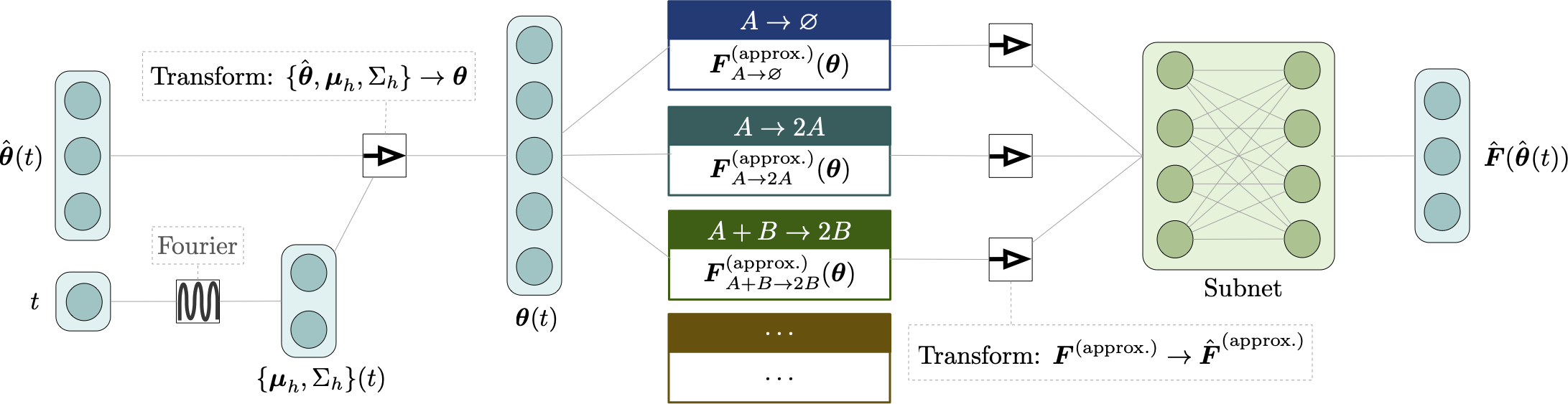
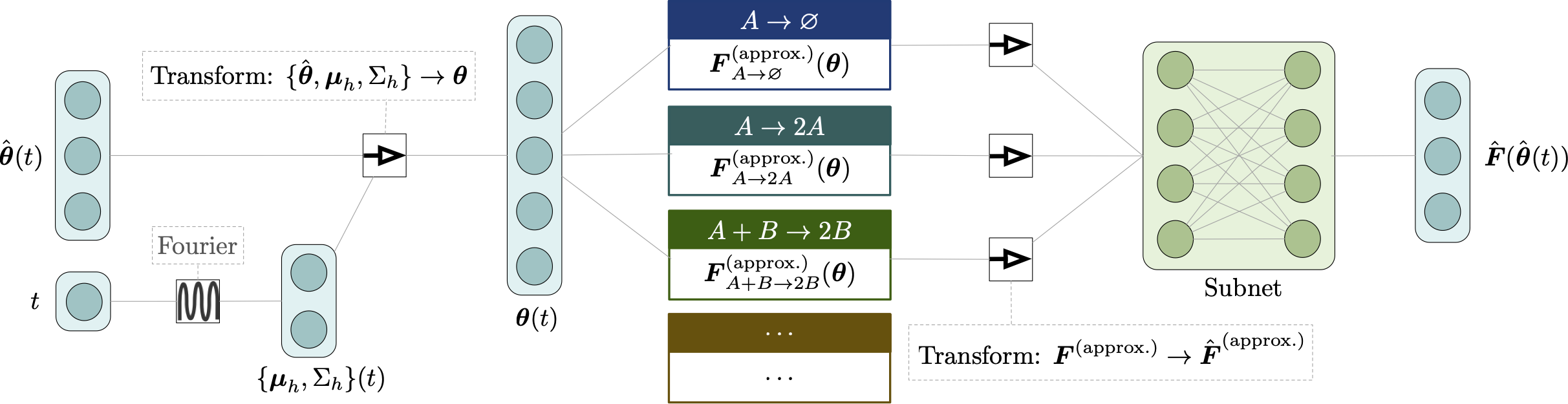
Deep learning moment closure approximations using dynamic Boltzmann distributions
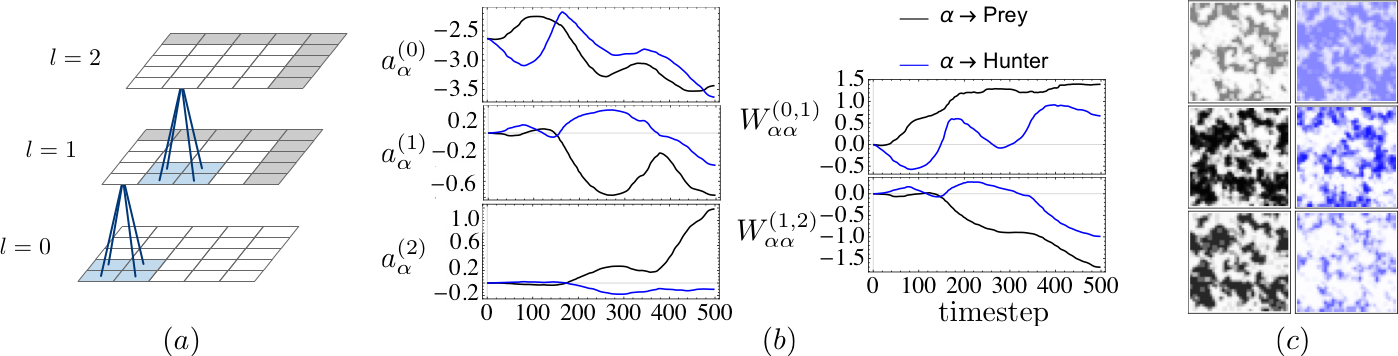
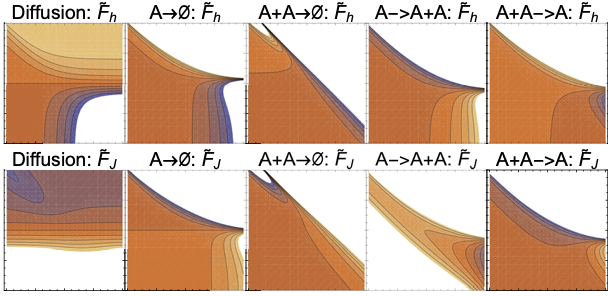
Learning moment closure in reaction-diffusion systems with spatial dynamic Boltzmann distributions
Learning dynamic Boltzmann distributions as reduced models of spatial chemical kinetics
Related work
- Graph-constrained Correlation Dynamics (GCCD)
Software
Various codes attached to papers (see above).
TensorFlow Python package for physics-based dynamic PCA
C++ library for Dynamic Boltzmann Machines for Lattice Chemical Kinetics
C++ library for Gillespie simulations on a lattice
C++ library for Gillespie simulations
Python package to sample pairs of particles from discrete Gaussians